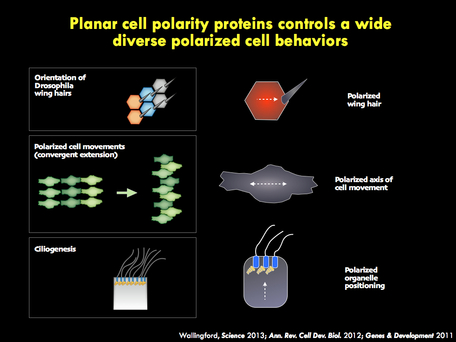
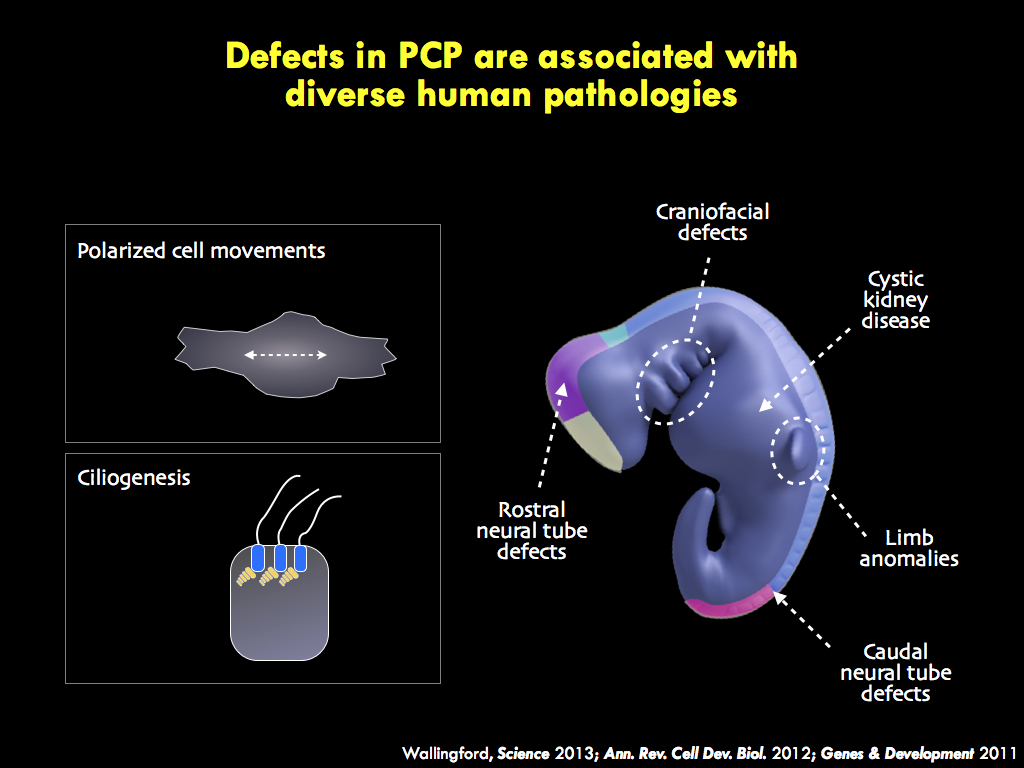
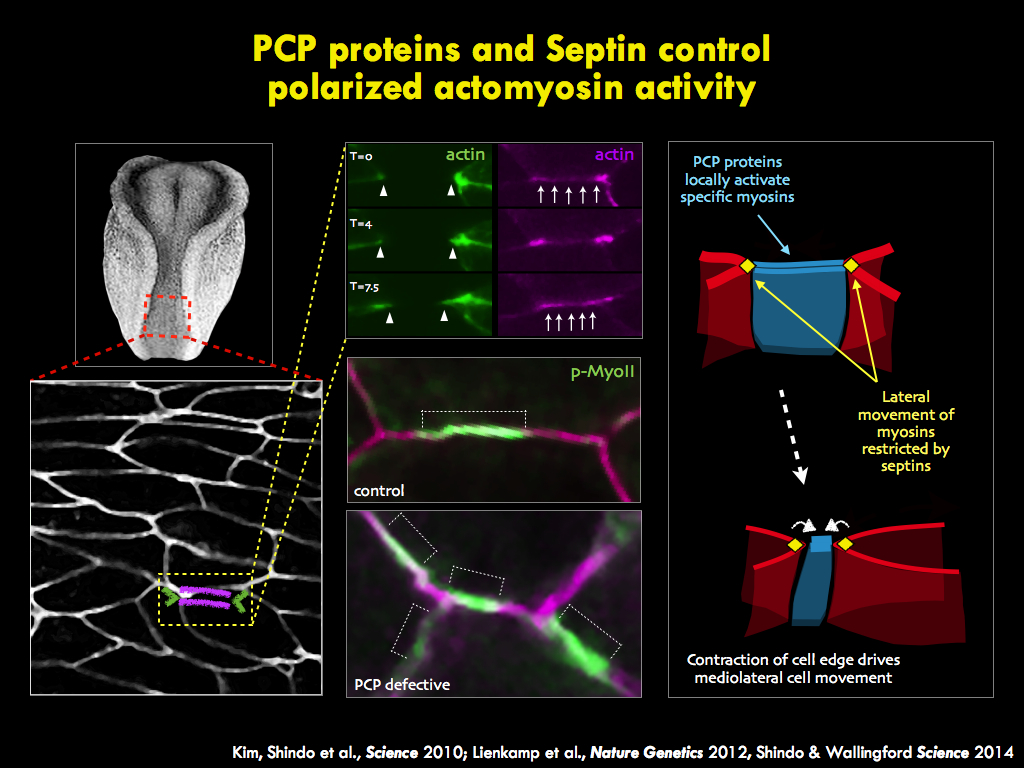
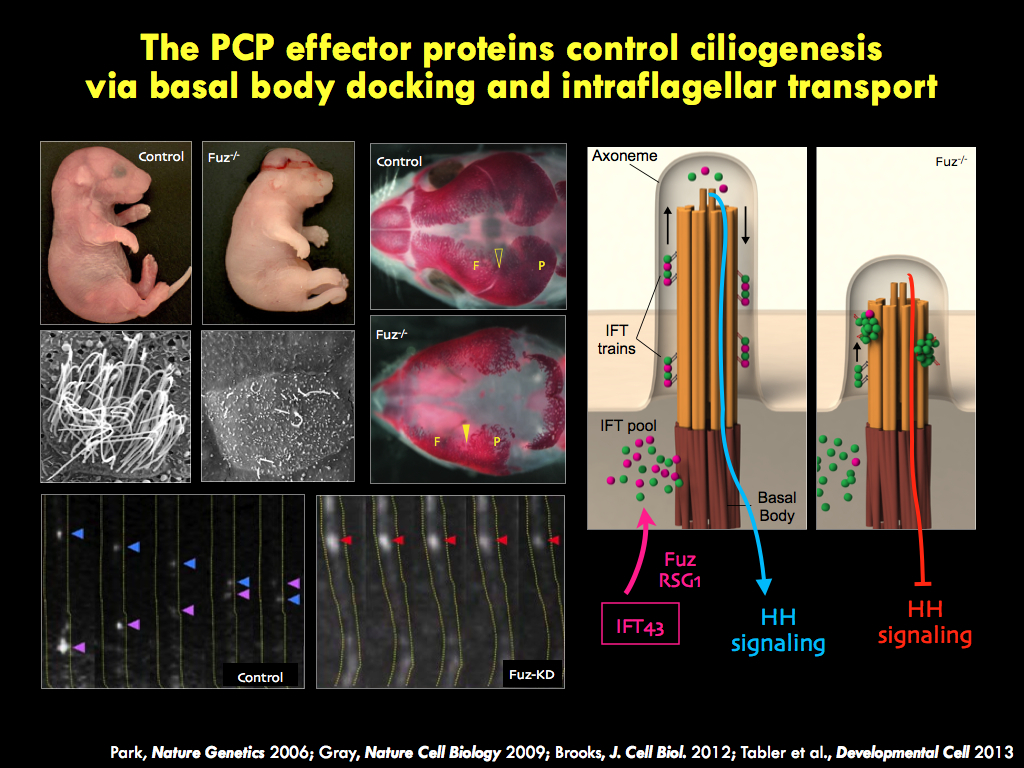
We seek to understand the molecular and cell biology of embryonic morphogenesis, with an eye toward
understanding the genetics of human birth defects such as spina bifida and ciliopathy.
understanding the genetics of human birth defects such as spina bifida and ciliopathy.
|
Why Xenopus? Coupled to a recent explosion of tools for genomic and proteomic analysis, the large cell size and externally developing embryos make Xenopus an ideal model for in vivo systems-level analysis of cell behavior in developing vertebrate embryos.
|
Why mouse? We insist that developmental cell biology must be pursued in vivo, so the mouse is a more challenging platform. But new tools are making live imaging more effective, and the unparalleled genetics in mice make the system essential to our work.
|
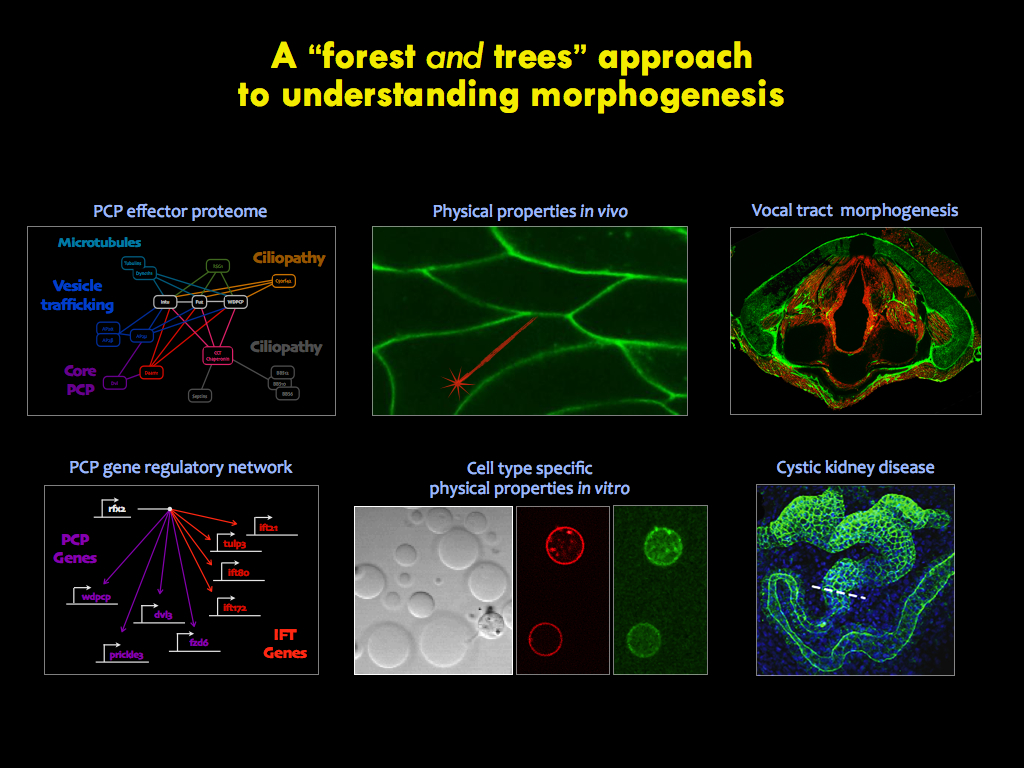
The panels below illustrate some of the major questions we address in the lab, some recent findings, and a snapshot of emerging themes in the lab.